We study the molecular mechanisms used by bacteria to display and utilize virulence factors. Our goal is to leverage this knowledge to discover novel antibiotics and to develop new tools for biotechnological applications. Our research is interdisciplinary, but primarily employs NMR, crystallography, biochemical and microbiological methods.
Pilus Assembly in Bacterial Pathogens
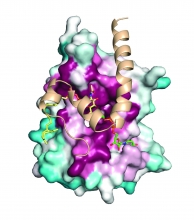
Bacterial pathogens display virulence factors that enable them to adhere to host tissues, resist phagocytic killing, invade host cells and acquire essential nutrients. We are studying how bacteria synthesize and display two types of virulence factors: (i) pili, proteinaceous fibers that project from the bacterial cell surface to mediate adhesion, and (ii) Wall Teichoic Acids (WTAs), highly...
Host-Pathogen Interactions: Heme-Iron Acquisition
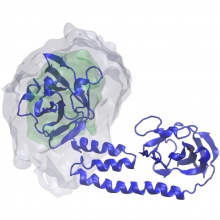
To successfully mount infections bacterial pathogens actively procure iron from their human host, which is extremely scarce because of nutritional immunity mechanisms. Hemoglobin within erythrocytes is an attractive source of iron, as it contains ~75-80% of the body’s total iron in the form of heme (iron-protoporphyrin IX). In ongoing research, we studying the molecular mechanisms used by Gram...
Chemical Biology: Antibiotic Discovery
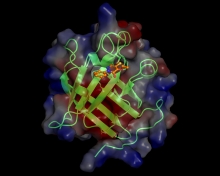
The emergence of antibiotic resistance bacteria is significant health concern. Our goal is to discover new anti-infective agents that can be used to combat infections caused by methicillin resistant Staphylococcus aureus (MRSA) and other multidrug resistant Gram-positive bacteria. Toward this objective, we are using high throughput screening approaches to identify small molecules that...
08/23 Christine Minor was awarded a CMB Fellowship! Congratulations!
-------------------------------------------------------------------------------------------------------------------------
07/23 Andrew Goring was awarded a UCLA Whitcome Fellowship!
-------------------------------------------------------------------------------------------------------------------------
05/23 Reece Pawlacyk joined our research group. Welcome!
-------------------------------------------------------------------------------------------------------------------------
08/22 Dr. Chris Sue was awarded his PhD in biochemistry! Congratulations!
-------------------------------------------------------------------------------------------------------------------------
06/22 Nikki Cheung and Jordan Ford were awarded CMB fellowships!
-------------------------------------------------------------------------------------------------------------------------
06/22 Nikki Cheung and Christine Minor joined our research group. Welcome!
-------------------------------------------------------------------------------------------------------------------------
05/22 Andrew Goring was awarded an Oral Health Predoctoral Fellowship!
-------------------------------------------------------------------------------------------------------------------------
04/22 Dr. Orlando Martinez was awarded his PhD in biochemistry!
-------------------------------------------------------------------------------------------------------------------------
03/21 Jordan Ford joined our research group. Welcome!
-------------------------------------------------------------------------------------------------------------------------
09/20 Dr. Scott McConnell was awarded his PhD in biochemistry!
-------------------------------------------------------------------------------------------------------------------------
06/20 Jess Soule was awarded a Chemistry-Biology Interface Training Grant!
-------------------------------------------------------------------------------------------------------------------------
06/20 Andrew Goring was awarded a Cellular and Molecular Biology Training Grant!
-------------------------------------------------------------------------------------------------------------------------