|
Role of Sumo-conjugation in Development
We have recently become interested in understanding
the biological role of a post-translational protein modification known
as Sumo-conjugation. Sumo is a small (~100 amino acid) protein with
about 20% homology to ubiquitin. Like ubiquitin, it is attached to other
proteins via an isopeptide linkage between its C-terminus and e-amino groups on the target proteins. While ubiquitylation
of proteins targets them for proteasomal degradation, the purpose of
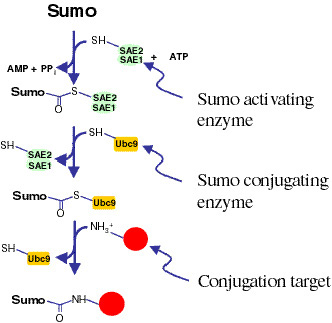
Sumo conjugation is not well understood. We have recently identified
the Drosophila genes encoding the enzymatic machinery responsible for
Sumo conjugation. This includes the genes encoding Sumo itself, the
Sumo-activating enzyme (SAE1/2), the Sumo-conjugating enzyme (Ubc9),
and the Sumo processing/deconjugating enzyme (Ulp1). Using a combination
of biochemical, molecular, and genetic analysis our goal is to illuminate
the biochemical and developmental roles of this process. Sumo-conjugation
may activate Dorsal in two ways. Our interest in Sumo-conjugation
was sparked by our discovery that Dorsal is a substrate for this modification.
Thus, our initial analysis of this pathway has focused on its interaction
with Dorsal (Bhaskar
et al., 2000; Bhaskar
et al., 2002). We have found that conjugation of Sumo to Dorsal stimulates this factor
in two ways. First, this modification appears to favor nuclear localization
of Dorsal. Since the conjugation machinery appears to reside in the
nucleus, we propose that the increase in nuclear localization may be
due to an effect of Sumo on nuclear retention as opposed to nuclear
import. Second, Sumo-conjugation appears to make Dorsal a more potent
transcriptional activator. To explore this effect, we mapped the lysine
in Dorsal to which Sumo is attached. Mutagenesis of this lysine resulted
in a constitutively activated form of the factor. These findings suggest
that, in its unmodified form, the Sumo-conjugation site may form part
of a docking surface for a negatively acting regulatory factor. We hypothesize
that either mutagenesis of the site or conjugation of Sumo to the site
results in displacement of the negatively acting factor and a consequent
increase in transcriptional activation. Sumo-conjugation
and the stress response. We suspect that conjugation of Sumo
to target proteins may be a stress response mechanism. The first line
of evidence in favor of this hypothesis derives from experiments in
which we examined the total level of Sumo-conjugates in Drosophila cells
before and after the cells have been subjected to environmental stresses
such as heat shock or high concentrations of hydrogen peroxide. In the
absence of stress, the majority of the Sumo is found in the unconjugated
form, but with increasing stress high molecular weight Sumo-conjugates
are formed depleting the pool of unconjugated Sumo. Apparently cellular
stress produces an intracellular signal that activates the conjugation
machinery. How
does Sumo-conjugation help cells cope with stress? So far our experiments
suggest two answers.
(1) We have observed that Sumo-conjugation is critically
required for the ability of fruit flies to mount a challenge to septic
injury (Bhaskar
et al., 2002). In particular, we have found that blocking Sumo-conjugation blocks the
activation of genes encoding antimicrobial peptides that usually occurs
in response microbial infection. This so-called innate immune response
is not normally considered to be a stress response in the same sense
that response to heat shock or hydrogen peroxide is considered a stress
response. However, our findings suggest that mechanistic similarities
may underlie these responses. (2)
We have immunopurified some of the high molecular weight conjugates
that appear in response to stress and identified them by mass spectroscopy.
Two of the three conjugates that we have identified are aminoacyl-tRNA
synthetases. Further analysis suggests that conjugation of Sumo to these
enzymes may facilitate the formation of a high molecular weight aminacyl-tRNA
synthetase complex that may coordinate tRNA charging. Thus, perhaps
Sumo-conjugation allows cells to cope with stress by activating tRNA
aminoacylation. This may help to replenish stores of aminoacyl-tRNAs
depleted by the synthesis of stress response proteins. Multiple
roles for a Sumo processing/deconjugating enzyme. Based on homology
to a gene encoding a yeast Sumo-deconjugating enzyme, we previously
identified a Drosophila gene (Ulp1) that encodes such an enzyme. We
found that, in addition to catalyzing the cleavage of isopeptide linkages
in Sumo-conjugated proteins, this enzyme can also cleave off the extension
found at the C-terminal end of the primary translation product of the
gene encoding Sumo. This cleavage process is required for the normal
maturation of Sumo. Thus, the enzyme appears to play both positive and
negative roles in the formation of Sumo-conjugates. This could explain
the biphasic response curve we see when we assay the activity of Dorsal,
a transcription factor that is stimulated by Sumo-conjugation, as a
function of Ulp1 concentration. Our
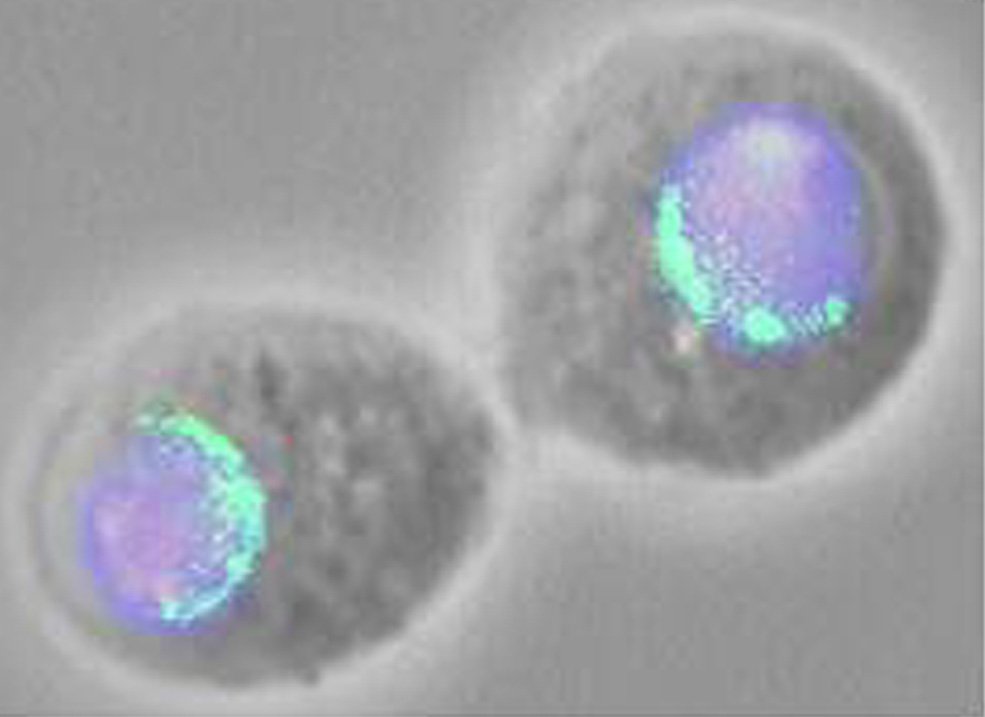
recent studies suggest yet another function for Ulp1. We expressed a
Ulp1-GFP fusion protein in Drosophila tissue culture cells and then
localized the protein by fluorescence imaging. We found that the fusion
protein is localized to multiple discrete foci in the nuclear membrane,
perhaps representing nuclear pore complexes. This finding suggests that
as a component of the nuclear pore complex, Ulp1 may play a role in
the nuclear import of Sumo and/or Sumo-conjugates. To test this idea,
we have examined the effect of knocking out Ulp1 activity on the subcellular
localization of Sumo. We find that that in the presence of functional
Ulp1 Sumo is largely nuclear, while in the absence of Ulp1 most of the
Sumo relocalizes to the cytoplasm. |